pISSN : 3058-423X eISSN: 3058-4302
Open Access, Peer-reviewed
pISSN : 3058-423X eISSN: 3058-4302
Open Access, Peer-reviewed
Mitthun ANK,Smiline Girija A.S,Vijayashree Priyadharsini J,Shoba Gunasekaran,Kannika Parameshwari Kannan
10.17966/JMI.2025.30.1.25 Epub 2025 April 01
Abstract
Background: Secreted aspartyl proteinases (SAPs) contribute to the pathogenicity of Candida albicans infections by aiding in tissue invasion and immune evasion.
Objective: This study aimed to identify SAP genes in clinical isolates of C. albicans and assess the antifungal potential of Aegle marmelos (also known as Bengal quince, golden apple, Japanese bitter orange, stone apple, or wood apple) using in vitro and in silico methods.
Methods: Carious scrapings were collected from patients with deep root caries. Standard microbiological techniques were employed for the phenotypic identification of C. albicans, and the presence of SAP genes was confirmed by PCR. A methanolic extract of A. marmelos was prepared and evaluated for antifungal activity against C. albicans isolates. In addition, in silico docking studies were conducted on six known bioactive compounds from A. marmelos to predict inhibitory effects on C. albicans SAPs.
Results: In total, 13 of 20 clinical samples (65.0%) tested positive for C. albicans, all of which were susceptible to growth suppression by the methanolic extract of A. marmelos. The furocourmarin imperatorin exhibited the highest binding affinity to SAP protein in silico (docking energy of -7.45 kcal/mol), likely by forming multiple hydrogen bonds, and showed no violations in molecular interactions.
Conclusion: Imperatonin may inhibit pathological C. albicans growth by inhibiting SAPs. Further experimental studies are warranted to confirm imperatorin toxicity, antifungal efficacy, and therapeutic potential.
Keywords
Aegle marmelos Candida albicans Health Imperatorin SAP protein
Candida albicans is a common opportunistic fungal patho- gen that resides in the oral and gastrointestinal tracts of 40~60% of healthy adults1. While generally a harmless com- mensal organism, C. albicans can transition to a pathogenic state in individuals with weakened immune systems, resulting in a fungal infection termed candidiasis2. It is often referred to as a yeast-like fungus because of its ability to form pseudo- hyphae3, so these infections are commonly referred to as "yeast infections".
Among Candida species, C. albicans is frequently used as a model organism to study fungal pathogens and to screen compounds for antifungal activity. Numerous virulence factors have been identified that promote candidiasis, including a family of at least 10 secreted aspartic proteases (SAPs)4. Furthermore, SAPs 1-3 were reported to exacerbate C. albicans-induced damage of reconstituted human epithelium in a model of human oral candidiasis and oropharyngeal candidiasis in individuals with HIV infection5,6. The genes encoding SAP1 and SAP2 are also frequently detected in vaginal candidiasis7. The cell surface-associated form SAP9 was demonstrated to efficiently and rapidly degrade the anti- microbial peptide histatin 5 produced by oral epithelial cells, thereby reducing endogenous resistance against C. albicans. In addition, SAP10 and SAP9 have been characterized as glycosylphosphatidylinositol (GPI)-anchored proteases that target proteins essential for cellular processes and host-pathogen interactions8. Thus, SAPs are considered essential virulence factors for C. albicans adhesion and invasion of epithelial cells and thus ultimately for tissue damage, but the precise contributions of individual SAP variants are not well described. Further, the host conditions conferring vulnerability and resistance to C. albicans infection are not well understood, although it has been shown that SAP variant activities are differentially sensitive to host tissue pH.
Numerous antifungal drugs are available to treated candid- iasis, but drug resistance among candidal strains has emerged as a major clinical problem in recent decades, necessitating the development or identification of alternative therapies. Many plants and herbs demonstrate antimicrobial and anti- fungal bioactivities. For instance, Aegle marmelos (commonly known as Bael) is a deciduous shrub or small to medium-sized tree with multiple documented bioactive properties9, which have been attributed to phytoconstituents such as marmenol, marmin, marmelosin, marmelide, psoralen, alloimperatorin, rutaretin, scopoletin, aegelin, marmelin, fagarine, anhydro- marmelin, and limonene10. Extensive experimental and clinical research has shown that Aegle marmelos processes anti- diarrheal, antimicrobial, antiviral, radioprotective, anticancer, chemopreventive, antipyretic, ulcer-healing, antigenotoxic, diuretic, antifertility, and anti-inflammatory effects11. The present study evaluated the expression of SAP genes in C. albicans samples from patients with root caries and the inhibitory effects of methanolic crude extracts from A. marmelos on C. albicans growth. In addition, an in silico analysis was con- ducted to investigate the drug-ligand interactions between SAP and six known bioactive compounds derived from A. marmelos, coumarin, xanthotoxol, imperatorin, aegeline, marmeline, and fluconazole.
1. Study Setting and preliminary identification of Candida albicans
This study was conducted at the Department of Micro- biology of Saveetha Dental College and Hospitals from April to June 2022. All protocols were approved by the institutional ethics review board (SRB/SDC/UG-2044/21/MICRO/063; IHEC/SDC/UG-2044/21/MICRO/604) and the study was con- ducted according to the tenets of the Declaration of Helsinki (1975 and subsequent amendments). Participants provided informed written consent prior to testing. Carious scrapings were collected from 20 patients with root caries as assessed by an endodontist. The samples were collected in sterile Sabouraud dextrose broth and promptly transported to the microbiology laboratory for processing. Samples were then inoculated on sterile Sabouraud dextrose agar (SDA) and incubated at 37℃ for 24 h. After incubation, colonies were identified based on their morphology and Gram staining. In addition, cultures were transferred to sterile Himedia Differen- tial Hicrome Agar for the rapid identification of C. albicans by.
2. Genotypic characterization of the sap gene in C. albicans
Fresh C. albicans cultures were cultured on Sabouraud dextrose agar (SDA) and incubated at 37℃ for 48 h. Genomic DNA was then extracted from isolates using the Qiagen kit according to the manufacturer's instructions. The SAP genes sap4 to sap7 were detected by PCR using a reaction mixture composed of 7.8 μl of 2× master mix (Taraka, Japan), 5.6 μl of double-distilled water, and 1.6 μl of the target primers (Table 1) [Eurofins Genomic India Pvt Ltd, Bangalore], and 35 amplification cycles with an annealing temperature of 58℃. The amplicons were then visualized by electrophoresis on a 1.5% agarose gel containing ethidium bromide and captured using a gel documentation system. Amplicon size was estimated using a 1.5-Kbp DNA ladder12.
|
Gene target |
Primers |
Annealing temperature |
Amplicon size |
|
sap4 |
5'-CATTCATTCCTTTAATACCGACTATC-3' |
58℃ |
156 |
|
5'-GGTAACAAACCCTGTAGATCTTTTAAC-3' |
|||
|
sap5 |
5'-CATTCATTCCTTTAATACCGACTATC-3' |
58℃ |
181 |
|
5'-GGTAACAAACCCTGTAGATCTTTTAAC-3' |
|||
|
sap6 |
5'-CATTCATTCCTTTAATACCGACTATC-3' |
58℃ |
206 |
|
5'-GGTAACAAACCCTGTAGATCTTTTAAC-3' |
|||
|
sap7 |
5'-GAAATGCAAAGAGTATTAGAGTTATTAC-3' |
58℃ |
196 |
|
5'-GAATGATTTGGTTTACATCATCTTCAACTG-3' |
3. Preparation of the A. marmelos extract
Fresh A. marmelos leaves were thoroughly washed three times with sterile distilled water to eliminate any impurities and external contaminants, dried in the shade, and ground into a coarse powder using a mechanical grinder. The powder was stored in a sterile container for later extraction. To prepare the crude extract, 10 g of the powdered leaves was mixed with 100 ml of methanol and incubated at room temperature with occasional agitation for one week. The crude extract was filtered through Whatman No. 1 filter paper and concentrated by evaporation in a sterile Petri dish. The final extract was stored at 4℃ for subsequent in vitro studies.
4. Antifungal bioassay
A 20-mg sample of A. marmelos extract was mixed with 1 ml of dimethyl sulfoxide (DMSO) and vortexed. Diluted samples of 5 and 10 mg/ml were prepared by adding more DMSO. The antifungal bioassay was conducted using the disc diffusion method. Briefly, C. albicans suspensions were pre- pared in sterile saline and adjusted to match the turbidity of a 0.5 McFarland standard (1.5 × 108 CFU/ml) for reproducibility. Lawn cultures were prepared by evenly spreading the adjusted suspension over the surface of sterile Sabouraud dextrose agar plates, and wells were created using an agar cutter. A 50 μl volume of extract (20, 10, and 5 mg/ml) or 50 μl DMSO (negative control to account for any potential antifungal properties of DMSO) was added to the respective wells. The plates were incubated for 48 h at 37℃, followed by measures of the zone of inhibition to assess the antifungal activity of the applied extract. The assay was repeated three times at each dose with independently seeded cultures, and the mean zone diameter calculated.
5. Retrieval of SAP protein structure
The structure of C. albicans SAP was retrieved from the Universal Protein Resource (UNIPROT) Database (ID 2QZW-A-Chain, https://www.uniprot.org/). Water molecules were removed and hydrogen atoms added using AutoDock, and the final three-dimensional structure was visualized using the Raster of a Molecule server13.
6. Preparation and optimization of the ligands
ChemSketch software was employed to determine the structures of the bioactive A. marmelos derivatives, coumarin, xanthotoxol, imperatorin, aegeline, marmeline, and fluco- nazole14. Structures were then optimized using an open-source molecular converter program and results saved in Protein Data Bank (PDB) format (.mol).
7. Drug likeness assessment by Molinspiration molecular parameters
The drug-likeness of the compounds was assessed by Lipinski's rule of five and by determining the number of hydrogen bond acceptors and donors (both of which are linked to bioavailability and membrane permeability), the partition coefficient (logP), and molecular weight using the Molinspiration tool (molinspiration.com). In addition, the ab- sorption, distribution, metabolism, and elimination (ADME) properties of each bioactive compound were analyzed using the ADME tool15.
8. Docking analysis and visualization of the drug ligand interactions
The interactions between SAP and the selected bioactive compounds derived from A. marmelos were simulated by docking analysis using the AutoDock tool and Discovery Studio Visualizer16. Docking score evaluations, affinities, molecular dynamics, and simulation energies were calculated to compare the relative stabilities of drug-ligand interactions.
1. Characterization of C. albicans in root caries scrapings
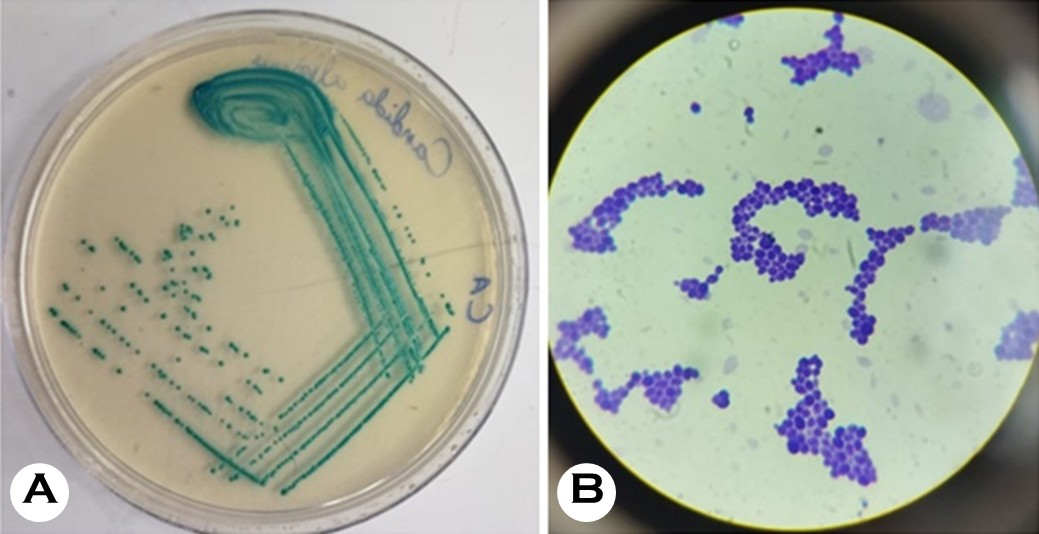
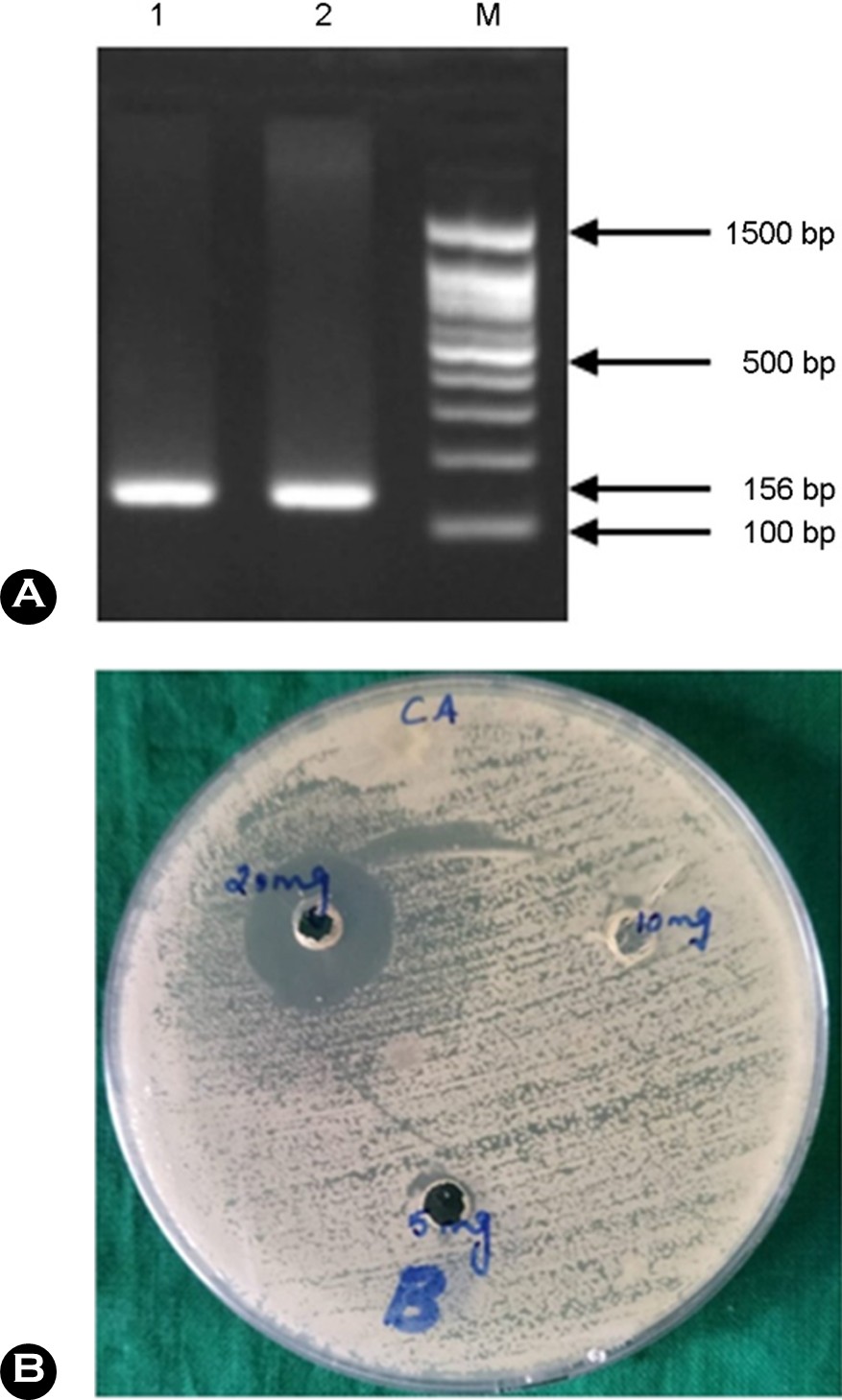
Of the 20 root caries scraping samples, C. albicans was detected in 13 (65.0%) as evidenced by the characteristic greenish-blue colonies on Hicrome Candida Differentiation Agar (Fig. 1A) and the Gram-positive staining, which revealed budding, oval-shaped cells (Fig. 1B). Among these 13 samples, three (23.07%) tested positive for the sap4 gene by PCR amplification, which yielded the predicted amplicon size of 156 bp (Fig. 2A).
2. Antifungal effect of the A. marmelos extract
The methanolic extraction procedure yielded 34 mg/ml from 100 g of dried A. marmelos leaf powder. Disk diffusion assays demonstrated substantial antifungal activity against all tested isolates at the 20 mg dose, with a zone of inhibition measuring 19 ± 1.5 mm, while no antifungal activity was observed at 10 and 5 mg doses (Fig. 2B). No zone of inhibition was observed around the wells containing 50 μl DMSO, confirming that the solvent did not substantially interfere with or promote the antifungal activity of the extract.
3. C. albicans SAP structure retrieval
The Fast-All sequence of the C. albicans sap gene was retrieved from the Universal Protein Resource (UNIPROT) database using the sequence ID 2QZW-A-Chain. The three-dimensional structure of SAP was visualized using the RAMSOL tool with default visualization parameters, including wire- frame view and coloring by atom type. The structure was manipulated to highlight the key residues involved in ligand-binding interactions.
4. Retrieval of A. marmelos bioactive ligand structures
Structures of the selected A. marmelos bioactive ligands coumarin, xanthotoxol, imperatorin, aegeline, marmeline, and fluconazole were drawn and optimized using ACD Chem- Sketch. After the initial 2D structure construction, geometric optimization was conducted using the MMFF94 (Merck Molecular Force Field) application to minimize the energy of the molecules. The optimized structures were then exported in PDB format, and an OpenBabel molecular converter was used to convert these structures into a compatible format for docking simulations. Table 2 displays the ligands from A. marmelos together with the simplified molecular input line entry system (SMILES) format.
|
Compound |
SMILES |
Mol. |
Mol |
Pubchem |
|
Coumarin |
C1=CC=C2C(=C1)C=CC(=O)O2 |
C9H6O2 |
146.14 |
323 |
|
Xanthotoxol |
C1=CC(=O)OC2=C(C3=C(C=CO3)C=C21)O |
C11H6O4 |
202.16 |
65090 |
|
Imperatorin |
CC(=CCOC1=C2C(=CC3=C1OC=C3)C=CC(=O)O2)C |
C16H14O4 |
270.28 |
10212 |
|
Aegeline |
COC1=CC=C(C=C1)C(CNC(=O)/C=C/C2=CC=CC=C2)O |
C18H19NO3 |
297.3 |
15558419 |
|
Marmeline |
CC(=CCOC1=CC=C(C=C1)C(CNC(=O)/C=C\C2=CC=CC=C2)O)C |
C22H25NO3 |
351.4 |
131750977 |
|
Erythromycin |
CC[C@@H]1[C@@]([C@@H]([C@H](C(=O)[C@@H](C[C@@]([C@@H]([C@H]([C@@H]([C@H](C(=O)O1)C)O[C@H]2C[C@@]([C@H]([C@@H](O2)C)O)(C)OC)C)O[C@H]3[C@@H]([C@H](C[C@H](O3)C)N(C)C)O)(C)O)C)C)O)(C)O |
C37H67NO13 |
733.9 |
12560 |
|
SMILES: Simplified Molecular Input Line
Entry System |
||||
5. Molinspiration assessment toward drug likeness
The drug-likeness of the selected compounds was assessed based on key molecular properties using the Molinspiration tool. Lipinski's rule of five was then applied to ensure that all compounds adhered to the guidelines without violation. Molecular weight, partition coefficient (logP), and numbers of hydrogen bond acceptors, hydrogen bond donors, and rotatable bonds were also assessed. Ion channel modulation, G protein coupled receptor ligand, nuclear receptor ligand, kinase inhibitor, enzyme inhibition, and protease inhibition calculations were used to derive bioactivity scores for each selected compound against the SAP protein of C. albicans (Table 3). All compounds showed zero violations. The bio- activity scores and properties of the selected compounds are shown in Table 4.
|
Compound |
Hydrogen bond |
Hydrogen bond |
miLogP |
Rotatable |
Violations (number) |
TPSA |
Mol. |
Atoms |
|
Coumarin |
0 |
2 |
2.01 |
0 |
0 |
30.21 |
128.59 |
11 |
|
Xanthotoxol |
1 |
4 |
2.00 |
0 |
0 |
63.58 |
162.16 |
15 |
|
Imperatorin |
0 |
4 |
3.95 |
3 |
0 |
52.59 |
240.47 |
20 |
|
Aegeline |
2 |
4 |
2.64 |
6 |
0 |
58.56 |
281.45 |
22 |
|
Marmeline |
2 |
4 |
4.32 |
8 |
0 |
58.56 |
342.23 |
26 |
|
Fluconazole |
1 |
7 |
-0.12 |
5 |
0 |
81.66 |
248.96 |
22 |
|
Note:
Drug likeliness scores > 0.3. TPSA:
topological polar surface area |
||||||||
|
EfbA |
H- Bonds |
Binding (kJ) |
Ki |
Ligand |
Inter- |
vdW + |
Electro- |
Torsional |
Total |
|
Coumarin |
1 |
-5.9 |
47.36 |
-0.54 |
-5.9 |
-5.81 |
-0.09 |
0.0 |
0.0 |
|
Xanthotoxol |
1 |
-6.27 |
25.16 |
-0.42 |
-6.57 |
-6.57 |
-0.07 |
0.3 |
-0.39 |
|
Imperatorin |
2 |
-7.45 |
3.44 |
-0.37 |
-8.35 |
-8.23 |
-0.12 |
0.89 |
-0.54 |
|
Aegeline |
2 |
-6.82 |
9.94 |
-0.31 |
-8.91 |
-8.42 |
-0.49 |
2.09 |
-0.68 |
|
Marmeline |
2 |
-6.36 |
21.91 |
-0.24 |
-9.04 |
-8.77 |
-0.27 |
2.68 |
-1.87 |
|
Fluconazole |
2 |
-4.86 |
274.92 |
-0.22 |
-6.65 |
-6.55 |
-0.09 |
1.79 |
-1.5 |
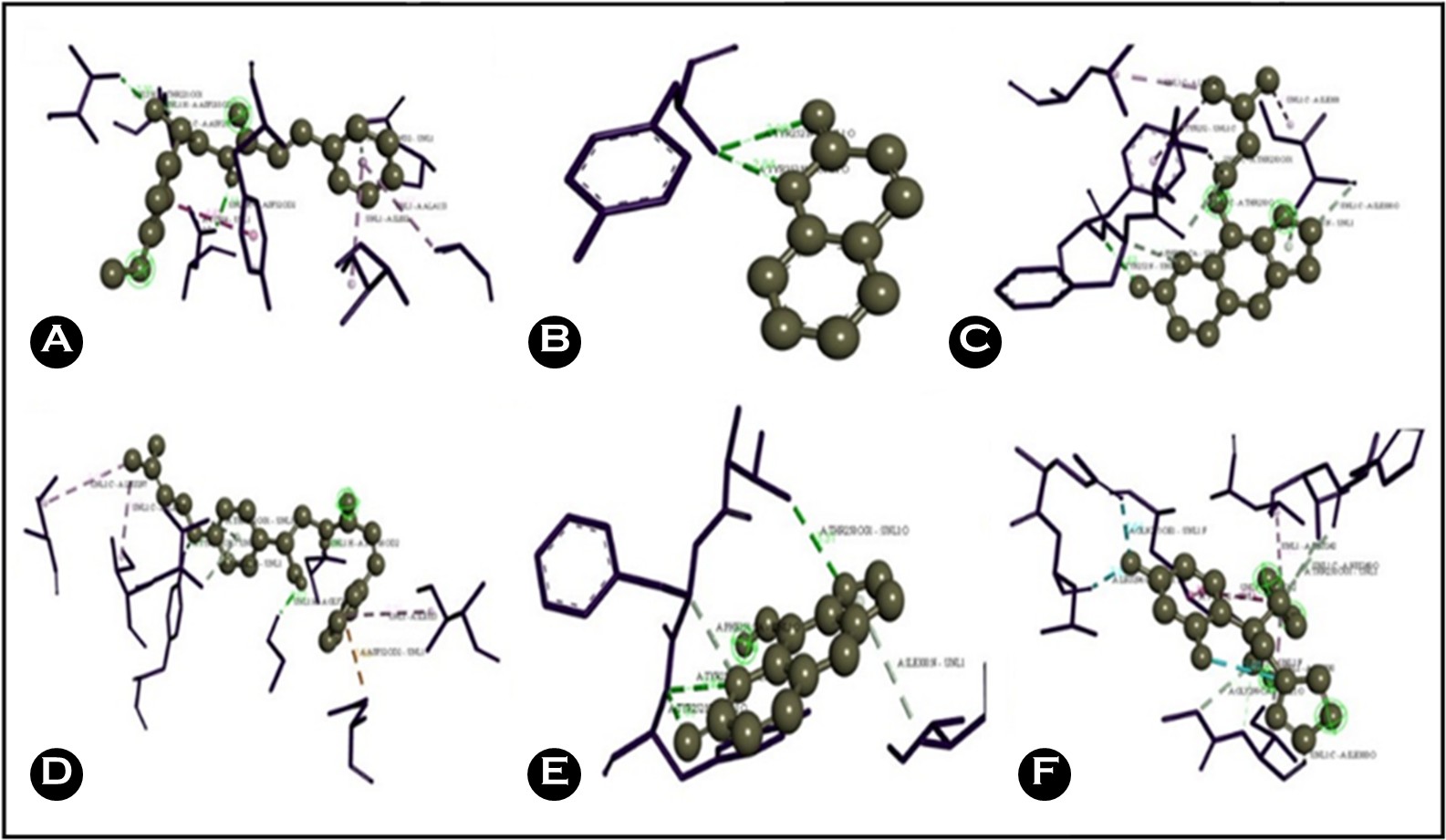
6. Docking results of the interactions between sap and bio-compounds from A. marmelos
The interactions between SAP and the aforementioned bioactive compounds from A. marmelos were then examined in silico using Discovery Studio Visualizer (Fig. 3). The SAP protein structure was prepared by removing water molecules and adding hydrogen atoms according to the AutoDock preparation protocol. Ligand structures were imported in compatible formats (.mol) for the docking simulations. Using AutoDock Vina, the best-docked pose with the lowest binding energy was selected for further analysis. Atomic interactions, including hydrogen bonds, van der Waals interactions, π-π stacking, and alkyl/π-alkyl interactions, were visualized using Discovery Studio, and the binding energy for each ligand-protein interaction was calculated. Among the compounds tested, imperatorin exhibited the most favorable binding energy and additional π-sulfur interactions (Table 5), suggest- ing that imperatorin is a major contributor to SAP inhibition and C. albicans growth suppression.
|
EfbA docking |
H-bonds |
Van der |
pi-sigma, |
Alkyl and |
pi-pi T-shaped |
Carbon- |
|
Coumarin |
1 |
- |
- |
- |
- |
7 |
|
Xanthotoxol |
3 |
3 |
- |
1 |
1 |
2 |
|
Imperatorin |
4 |
4 |
- |
1 |
- |
3 |
|
Aegeline |
3 |
10 |
- |
2 |
- |
1 |
|
Marmeline |
3 |
5 |
1 |
2 |
- |
1 |
|
Fluconazole |
2 |
5 |
- |
- |
- |
1 |
The polymorphic fungus C. albicans is a part of the normal human microbiome in most individuals17. However, C. albicans is an opportunistic pathogen capable of causing a variety of infections under certain conditions, ranging from mild skin conditions to severe systemic diseases. Several virulence factors have been identified that contribute to the pathogenesis of this yeast-like fungus and greatly enhance its ability to cause infection18. Most of these virulent factors assist in infection by adhering to host cells, enhancing host cell invasion through hydrolase activity, aiding in the transition of the yeast-to-hyphal stage, acting as biosensors, promoting the formation of biofilms, and (or) by facilitating phenotype switching19. Although C. albicans overgrowth is commonly associated with oro-dental infections such as oral candidiasis, it can also contribute to the development of root caries, particularly in immunocompromised individuals20. Moreover, oral candidiasis in immunocompromised individuals can progress to a recal- citrant infection throughout the oropharynx21. The over- growth of C. albicans in the oral cavity may also promote the formation of biofilms that increase the severity of other infections and conditions, such as root caries caused by Streptococcus mutans and other bacteria22, especially under conditions of dysbiosis from immunosuppression or poor oral hygiene23. The capacity of C. albicans to exacerbate root caries stems from its ability to adhere to tooth surfaces and produce degradative hydrolytic enzymes24. Additionally, C. albicans can shift from a commensal to a pathogenic state, further enhancing candidiasis risk and caries progression25. Understanding the interplay between C. albicans and other microorganisms in the oral cavity is thus essential for devel- oping targeted therapeutic strategies against candidiasis and root caries.
In the current study, C. albicans was identified in the majority of scrapping samples from root caries (65%), consistent with contributions to caries development, and sap4 was detected in a fraction of these samples. Recombinant sap1, 2, 3, and 6 can induce the release of proinflammatory cytokines from human monocytes, and sap4 to sap6 (but not sap1 to sap3) also contribute to the survival and escape from macrophage-induced destruction8. In addition to the sap-mediated viru- lence mechanisms, strains resistant to common antifungal agents have emerged, presenting a challenge to treatment using conventional therapies. For instance, several studies have reported strains of C. albicans resistant to amphotericin B and fluconazole26. In the current study, we selected sap4 to sap7 for PCR detection based on documented involvement in virulence and pathogenicity. Indeed, previous studies have highlighted the significant roles of these gene products in mediating host-pathogen interactions, tissue invasion, and survival during infection. Thus, these variants are likely among the predominant virulence factors in the pathogenesis of C. albicans infections27.
Therefore, this study tested the efficacy of a methanolic extract derived from A. marmelos, a medicinal plant with a variety of documented bioactivities and a well-characterized bioactive compound profile. Many such studies have empha- sized the therapeutic potential of medicinal plants due to well-documented safety profiles and global availability28,29, and A. marmelos leaf has shown notable efficacy against other dermatophytes30 and pathogens31. In this study as well, assessment using the disc diffusion method demonstrated substantial C. albicans growth inhibition at a dose of 20 mg, suggesting therapeutic utility against oral infections.
In silico analysis further verified that individual phyto- chemicals contained in A. marmelos extract bound with high affinity to SAPs. Of the compounds investigated, impera- torin demonstrated the highest affinity (binding energy of -7.45 kcal/mol), followed by angelin (-6.82) and marmeline (-6.36). The highest number of hydrogen bonds was observed with imperatorin when compared to the fluconazole, which had only two hydrogen bonds. These findings are consistent with previous in silico-based assessments32,33 and reports of promising bioactivities of imperatorin and the other assayed compounds against systemic infections34. For instance, cou- marins are widely utilized in the treatment of prostate cancer, renal cell carcinoma, and leukemia, and also help mitigate the side effects of radiotherapy. Xanthotoxol, another furano- coumarin, is a key active component of Cnidium monnieri35. The extract of A. dahurica also contains imperatorin, which has sedative properties in addition to efficacy against head- aches, toothaches, colds, and rheumatism. Aegeline is often marketed for enhancing weight loss, athletic performance, and vitality, although there is inadequate scientific evidence to support these claims36. Marmeline, an organic phenol ether, also has potent bioactive properties37. Further investigation is warranted to assess the contributions of these individual compounds to C. albicans growth suppression as well as other antifungal activities.
This study demonstrated the antifungal potential of A. marmelos against C. albicans using a combination of in vitro and in silico approaches. However, the study has limitations. First, we did not test the relative efficacy of A. marmelos extract against clinical antifungal agents such as fluconazole. The small sample size (N = 20) also limits the statistical power of the results. Third, we detected only sap4 to sap7 by PCR. While the in silico docking analysis identified promising inter- actions between A. marmelos bioactive compounds and SAP proteins, we did not assess the direct antifungal effects of these individual phytochemicals. Furthermore, the potential cytotoxic effects of A. marmelos extracts on human cells were not evaluated, which is critical for safe therapeutic applications. To establish the clinical relevance of A. marmelos, future studies should include comparative analyses with standard antifungal agents, larger sample sizes, in vivo validations, and cytotoxicity assessments.
The prevalence of C. albicans in oro-dental infections is rising in both healthy and immunosuppressed individuals. The emergence of antifungal resistance and the frequent expression of SAP-like virulence factors by Candida are serious threats complicating treatment, necessitating alternative therapies. The present study supports the efficacy of imperatorin from A. marmelos as a suitable drug candidate for further preclinical and possible clinical evaluation.
References
1. Wagner AS, Vogel AK, Lumsdaine SW, Phillips EK, Willems HME, Peters BM, et al. Mucosal infection with unmasked Candida albicans cells impacts disease pro- gression in a host niche-specific manner. Infect Immun 2022;90:e0034222
Google Scholar
2. Kirsch DR. Progress in Candida albicans molecular biology. In Candida albicans: Cellular and molecular biology. Berlin, Heidelberg: Springer Berlin Heidelberg, 1991:198-213
Google Scholar
3. Beema Shafreen RM, Seema S, Alagu Lakshmi S, Srivathsan A, Tamilmuhilan K, Shrestha A, et al. In vitro and In vivo antibiofilm potential of eicosane against Candida albicans. Appl Biochem Biotechnol 2022;194: 4800-4816
Google Scholar
4. Wu T. Secreted aspartyl proteinases of Candida albicans with particular relevance to the oral cavity. HKU Theses Online (HKUTO) 1997
Google Scholar
5. White TC, Miyasaki SH, Agabian N. Three distinct secreted aspartyl proteinases in Candida albicans. J Bacteriol 1993; 175:6126-6133
Google Scholar
6. Wu T, Samaranayake LP. The expression of secreted aspartyl proteinases of Candida species in human whole saliva. J Med Microbiol 1999;48:711-720
Google Scholar
7. Research CM, Case Medical Research. A study of oral VT-1161 for the treatment of acute vaginal candidiasis (yeast infection) in patients with recurrent vaginal can- didiasis. Case Medical Research 2000
8. Borg-von Zepelin M, Beggah S, Boggian K, Sanglard D, Monod M. The expression of the secreted aspartyl pro- teinases Sap4 to Sap6 from Candida albicans in murine macrophages. Mol Microbiol 1998;28:543-554
Google Scholar
9. Tan B, Chiranthanut N, Chansakaow S, Sireeratawong S, Khonsung P, Nimlamool W, et al. Anti-inflammatory effects of Pikad Tri-phol-sa-mut-than remedy, consisting of dried fruits of Aegle marmelos (L.) Corrêa, Coriandrum sativum L., and Morinda citrifolia L. J Ethnopharmacol 2022;298:115639
Google Scholar
10. Kiratipaiboon C, Dasuni Wasana PW, Hasriadi, Sukrong S, Ruangrungsri N, Towiwat P. Herbal root extracts in Ben-Cha-Moon-Yai remedy attenuated pain-like behaviors and inflammation through the opioid and prostaglandin systems. J Ethnopharmacol 2022;290:115088
Google Scholar
11. Rahman S, Parvin R. Therapeutic potential of Aegle marmelos (L.)-An overview. Asian Pac J Trop Dis 2014;4: 71-77
Google Scholar
12. Safiya SN, Girija ASS, Priyadharsini VJ. Molecular detec- tion of secreted aspartyl proteinases (Saps) from dental isolates of Candida albicans and targeting with Psidium guajava biocompounds: An in vitro and in silico analysis. Cureus 2023;15:491-543
Google Scholar
13. Fraley AD, Odegaard KE, Vilchiz VH, Kang C, Lee C. Exploring protein structure with RasMol, a free online tool. Am Biol Teach 2022;84:369-373
Google Scholar
14. Kumar P, Alpana A, Dinesh C, Abhinav B. In silico screening and molecular docking of bioactive agents towards human coronavirus receptor. GSC Biol Pharm Sci 2020;11:132-140
Google Scholar
15. Yu H, Adedoyin A. ADME-Tox in drug discovery: integra- tion of experimental and computational technologies. Drug Discov Today 2003;8:852-861
Google Scholar
16. Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 2010;31:455-461
Google Scholar
17. Mackenzie DW. Candida albicans: The pathogenic fungus. J Clin Pathol 1990;43:439
Google Scholar
18. Cassone A. Chitin synthesis, dimorphism and virulence in the pathogenic fungus Candida albicans. In Chitin in nature and technology. Boston, MA: Springer US 1986: 71-78
Google Scholar
19. Bu QR, Bao MY, Yang Y, Wang TM, Wang CZ. Targeting virulence factors of Candida albicans with natural products. Foods 2022;11:2951
Google Scholar
20. Pavithra AS, Maiti S, Rajeshkumar S, Jessy P. Antimicrobial study of cast post and its surface modification with nanoparticle. J Adv Pharm Technol Res 2022;13:s416-420
Google Scholar
21. Karajacob AS, Azizan NB, Al-Maleki ARM, Goh JPE, Loke MF, Khor HM, et al. Candida species and oral mycobiota of patients clinically diagnosed with oral thrush. PLoS One 2023;18:e028043
Google Scholar
22. Salehi B, Kregiel D, Mahady G, Sharifi-Rad J, Martins N, Rodrigues CF. Management of Streptococcus mutans-Candida spp. Oral biofilms' infections: Paving the way for effective clinical interventions. J Clin Med 2020;9: 517
Google Scholar
23. Kashyap B, Padala SR, Kaur G, Kullaa A. Candida albicans induces oral microbial dysbiosis and promotes oral diseases. Microorganisms 2024;12:2138
Google Scholar
24. Du Q, Ren B, He J, Peng X, Guo Q, Zheng L, et al. Candida albicans promotes tooth decay by inducing oral microbial dysbiosis. ISME J 2021;15:894-908
Google Scholar
25. Jørgensen MR. Pathophysiological microenvironments in oral candidiasis. APMIS 2024;132:956-973
Google Scholar
26. Bhattacharjee P. Epidemiology and antifungal suscepti- bility of Candida species in a tertiary care hospital, Kolkata, India. Curr Med Mycol 2016;2:20-27
Google Scholar
27. Ranasinghe A, Girija AS, Priyadharsini JV. Targeting the secreted aspartic proteinase (SAP-1) associated with virulence in C. albicans by C. cassia bio-compounds: a computational approach. J Pharm Res Int 2020;32:75-86
Google Scholar
28. Panda P, Das B, Bhuyan GC, Rao MM. Therapeutic uses of Vilwa (Aegle marmelos) in Ayurveda with pharmaco- logical evidence. Research Journal of Pharmacology and Pharmacodynamics 2021;13:51-54
Google Scholar
29. Sarojini K, Arivarasu L, Smiline Girija AS. Herbal formula- tion: Review of efficacy, safety, and regulations. Int J Res Pharmac Sci 2020;11:1506-1510
30. Choudhary Y, Saxena A, Kumar Y, Kumar S, Pratap V. Phytochemistry, pharmacological and traditional uses of Aegle marmelos. UK Journal of Pharmaceutical and Biosciences 2017;5:27-33
Google Scholar
31. Baliga MS, Bhat HP, Joseph N, Fazal F. Phytochemistry and medicinal uses of the bael fruit (Aegle marmelos Correa): A concise review. Food Res Int 2011;44:1768-1775
Google Scholar
32. Girija AS, Duraipandiyan V, Suba KP, Hariprasad G, Raghuraman R. Assessment of the anti-HBV like pro- perty of BEHP-a phthalate derivative and its in silico characterization. BMC Infect Dis 2014;14:1
33. Balamithra S, Girija S, Priyadharsini JV. An in silico analysis of protein targeted by glycyrrhizin in common dental pathogens. J Pharm Res Int 2021;32;170-178
Google Scholar
34. Awdhesh Kumar Mishra R, Kodiveri Muthukaliannan G. In-silico and in-vitro study of novel antimicrobial pep- tide AM1 from Aegle marmelos against drug-resistant Staphylococcus aureus. Sci Rep 2024;14:25822
Google Scholar
35. Sun Y, Yang AWH, Lenon GB. Phytochemistry, ethnophar- macology, pharmacokinetics and toxicology of Cnidium monnieri (L.) Cusson. Int J Mol Sci 2020;21:1006
Google Scholar
36. Zhu C, Wang M, Guo J, Su SL, Yu G, Yang Y, et al. Angelica dahurica extracts attenuate CFA-induced inflam- matory pain via TRPV1 in mice. Evid Based Complement Alternat Med 2022;2022:4684830
Google Scholar
37. Monika S, Thirumal M, Kumar PR. Phytochemical and biological review of Aegle marmelos Linn. Future Sci OA 2023;9:FSO849
Google Scholar
Congratulatory MessageClick here!