pISSN : 3058-423X eISSN: 3058-4302
Open Access, Peer-reviewed
pISSN : 3058-423X eISSN: 3058-4302
Open Access, Peer-reviewed
Jeyanthi Lalitha S,Sujitha R K,Srinivas K
10.17966/JMI.2025.30.1.18 Epub 2025 April 01
Abstract
Bacterial persisters represent a unique subset of bacteria that endure antibiotic exposure and pose a significant challenge to clinical treatment. Despite extensive studies and reviews on antibiotic persistence, understanding its underlying mechanisms remains elusive. A common misconception conflates persistence with mechanisms, such as resistance, heteroresistance, tolerance, and immune evasion, which can result in diagnostic and therapeutic complications. Distinguishing these processes is important for improving the management of persistent infections and designing effective treatments. This review summarizes bacterial persistence mechanisms, focusing on phenotypic heterogeneity, stress responses, and adaptive processes. The phenomenon is different from resistance and tolerance because of genetic stability and population dynamics. Furthermore, the ability of persisters to revert to a susceptible phenotype following the cessation of antibiotic exposure highlights the importance of targeted therapeutic approaches. Common persisters, including ESKAPE pathogens, emphasize the clinical and economic burden posed by antibiotic persistence. This review aims to provide insight into persistence to inspire novel strategies for combating antibiotic-resistant infections and improving therapeutic outcomes.
Keywords
Antibiotic persistence Biofilms Chronic infections Dormancy ESKAPE pathogens Toxin-antitoxin systems
Antibiotics have revolutionized modern medicine by sig- nificantly reducing mortality from bacterial infections and transforming healthcare. Despite these advancements, their overall effectiveness is limited by two critical phenomena: resistance and persistence. Resistance involves genetic changes that enable bacterial proliferation in the presence of antibiotics, whereas persistence is a phenotypic adaptation that enables survival without genetic alterations. These differences are important because persistence complicates treatment regi- mens and results in recurrent infections and therapy failures.
Joseph Bigger first identified bacterial persisters in 1944 as a subpopulation capable of surviving antibiotic exposure despite the eradication of the majority of the bacterial popula- tion. These persisters, which remain dormant during treat- ment, can eventually repopulate and cause relapse after the cessation of therapy. The significance of bacterial persistence has garnered increased attention, as evidenced by the 2018 European Molecular Biology Organization workshop (Table 1), which brought together 121 investigators from 21 countries to discuss bacterial persistence and antimicrobial therapy. Therefore, understanding persistence is necessary for the development of effective treatment strategies, particularly because it is distinct from resistance, heteroresistance, and tolerance1-5.
|
Workshop |
EMBO |
|
Topic |
Bacterial persistence and
antimicrobial therapy6 |
|
Held
on |
10~14
June 2018 |
|
Held
at |
Ascona,
Switzerland |
|
Number
of investigators |
121 |
|
Number
of participants |
21
countries6 |
Persistence is a survival strategy used by bacteria to survive antibiotic exposure. Unlike resistance, which is heritable and affects the entire population, persistence is transient and limited to a subset of cells. It arises from phenotypic hetero- geneity and stress-induced dormancy, which enables bacteria to survive lethal conditions.
1. Toxin-Antitoxin (TA) modules
TA modules are genetic systems consisting of a stable toxin and a labile antitoxin. These modules are ubiquitous in bacteria and regulate dormancy. Under normal conditions, the antitoxin neutralizes the toxin's activity. During stress, such as nutrient deprivation or antibiotic exposure, the antitoxin is degraded, freeing the toxin to inhibit essential cellular functions. This induces dormancy, which enables bacteria to the antibiotic's effects and promotes survival6,7.
2. Stringent response
The stringent response is a stress-induced mechanism triggered by nutrient deprivation, oxidative stress, or antibiotic exposure. It involves the production of signaling molecules, such as (p)ppGpp, that suppress metabolic and growth-related processes. This response conserves energy and resources, which facilitates bacterial dormancy and persistence8,9.
3. Metabolic shifts
Persisters undergo metabolic reprogramming to survive adverse conditions. A hallmark of persistence is decreased ATP production, which limits the activity of the processes targeted by antibiotics. In addition, alterations in biosynthetic pathways further reduce susceptibility to antimicrobials. These metabolic shifts are key to the persistence phenomenon and represent targets for therapeutic intervention10,11.
4. Genome-Wide Insights
Advances in genome-wide studies and transcriptomic pro- filing have identified numerous genetic pathways associated with persistence. Mutations affecting energy metabolism, stress responses, and cell envelope integrity are associated with persister formation. These findings have paved the way for novel therapies designed to disrupt the persistence mechanisms12-14.
Understanding the fundamental differences between per- sistence and related phenomena is important for developing effective treatments in Table 2.
|
Characteristic |
Resistance |
Persistence |
Heteroresistance |
|
Genetic
basis |
Stable genetic mutations or acquired genes |
No genetic changes; |
Genetic variants in the |
|
Population
affected |
Entire population |
Small subpopulation |
Variable subpopulation |
|
Heritability |
Heritable |
Nonheritable |
Partially heritable |
|
Growth
in antibiotics |
Can grow and divide |
Cannot grow or divide |
The subset can grow |
|
Minimum
inhibitory |
Increased |
Unchanged |
Variable within the |
|
Reversibility |
Generally permanent |
Reversible upon antibiotic |
Partially reversible |
|
Treatment
impact |
Complete treatment failure |
Delayed treatment response |
Variable response |
|
Clinical
detection |
Standard susceptibility testing |
Specialized persistence assays |
Population analysis is profiling |
1. Resistance
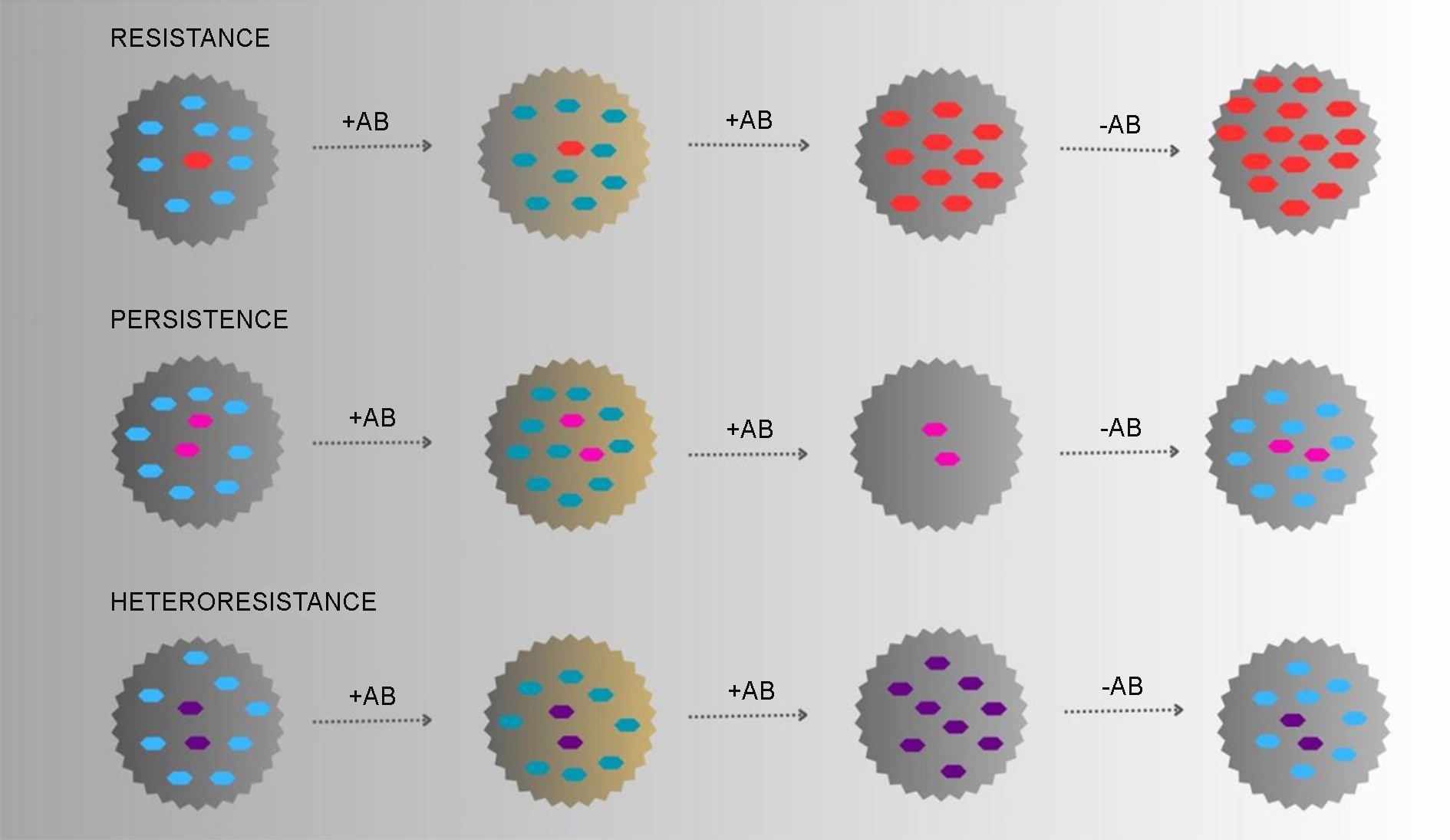
Resistance involves genetic mutations or horizontal gene transfer. This enables bacterial populations to grow and proliferate in the presence of antibiotics. Resistance is heritable and affects the entire population, whereas persistence is non-heritable and limited to a subpopulation of dormant cells34,35.
2. Heteroresistance
Heteroresistance refers to a transient subpopulation within a bacterial colony that exhibits higher minimum inhibitory concentrations compared with the majority. Unlike persistence, heteroresistant cells actively divide under specific conditions, whereas persisters remain dormant36,37.
A comparative study of the mechanisms of Resistance (A), Persistence (B), and Heteroresistance (C) is provided in Fig. 1.
3. Tolerance
Tolerance is the ability of a bacterial population to survive prolonged antibiotic exposure without experiencing genetic changes. Unlike persistence, which involves a small subpop- ulation, tolerance affects the entire population and delays bacterial killing rather than preventing it entirely38,39.
4. Biphasic killing curves
The hallmark of persistence is a biphasic killing curve, which shows an initial rapid decline in bacterial numbers resulting from the death of the susceptible population, followed by a plateau indicating survival of a nondividing persister sub- population40-42.
5. ESKAPE pathogens and persistence
The ESKAPE pathogens, Enterococcus faecium, Staphylo- coccus aureus, Klebsiella pneumoniae, Acinetobacter bau- mannii, Pseudomonas aeruginosa, and Enterobacter species, are bacteria known for causing multidrug-resistant and per- sistent infections. These pathogens pose significant clinical challenges because of their ability to survive aggressive treat- ments.
1) Enterococcus faecium (E. faecium)
E. faecium is a Gram-positive bacterium renowned for its resistance to vancomycin, which is a last-resort antibiotic. Its persistence is often linked to biofilm formation, which creates a protective niche and robust stress-response pathways that shield it from antibiotics and the host immune response. These mechanisms complicate treatment in hospital settings, particularly in immunocompromised patients15-17.
2) Staphylococcus aureus (S. aureus)
Methicillin-resistant S. aureus (MRSA) exemplifies persist- ence. MRSA strains utilize metabolic dormancy and biofilm formation to survive, rendering them particularly adept at evading antibiotics. These infections are notoriously difficult to eradicate, often resulting in chronic and recurrent cases18-21.
3) Klebsiella pneumoniae (K. pneumoniae)
K. pneumoniae is a major cause of nosocomial infections, particularly in intensive care units. It is known for producing carbapenemase that degrades carbapenems, which is a class of last-resort antibiotics. It utilizes biofilm formation and meta- bolic dormancy to promote persistence, further exacerbating its impact in healthcare settings22,23.
4) Acinetobacter baumannii (A. baumannii)
A. baumannii is frequently associated with infections in intensive care units, particularly in immunocompromised patients. Persistent strains evade standard treatments and immune responses through various mechanisms, such as bio- film formation and metabolic adaptation. These characteristics make A. baumannii a leading cause of hospital-acquired infections24-26.
5) Pseudomonas aeruginosa (P. aeruginosa)
P. aeruginosa is a model organism for studying persistence. Its intrinsic resistance mechanisms, which include efflux pumps and biofilm formation, enable it to survive hostile environ- ments, including the lungs of patients with cystic fibrosis. Chronic infections caused by P. aeruginosa are particularly challenging to treat because of persistence27-30.
6) Enterobacter species
Carbapenemase-producing Enterobacter species are a growing concern in clinical settings. These bacteria demon- strate persistence through biofilm formation and stress-induced dormancy, which complicates treatment and often results in recurrent infections31-33.
Addressing persistence requires innovative therapeutic approaches that target dormant cells, enhance antibiotic efficacy, and prevent recurrent infections.
1. Metabolic activation
Reactivating the metabolism of dormant cells renders them susceptible to antibiotics. Drugs that target metabolic pathways and stimulate ATP production have shown potential in experimental models and can eradicate persisters43,44.
2. Combination therapies
Combining bactericidal agents with anti-persistent com- pounds represents a synergistic approach to overcoming persistence. For example, pairing antibiotics with drugs that disrupt biofilms or inhibit stress-response pathways can en- hance treatment outcomes and reduce recurrence rates45-47.
3. Bacteriophage therapy
Bacteriophages, which are viruses that infect bacteria, are emerging as promising agents against persisters. They can penetrate biofilms and target dormant cells, which provides a potential adjunct to traditional antibiotics. Clinical trials evaluating phage therapy have yielded encouraging results48,49.
Accurate diagnostics are essential for detecting persistence in clinical settings. Emerging technologies, such as single-cell imaging, transcriptomics, and metabolomics, have improved our ability to identify persister populations and tailor treatments. Precision medicine that integrates host-pathogen interactions holds significant promise for mitigating persistence-driven infections50-52.
Bacterial persistence poses a significant challenge in the global fight against antibiotic-resistant infections. Based on their ability to survive treatment, persisters contribute to chronic and recurrent infections and impose substantial clinical and economic burdens. This review underscores the import- ance of multidisciplinary research to identify the mechanisms of persistence, differentiate them from related phenomena, and develop targeted therapeutic strategies. Addressing persistence is necessary for safeguarding antibiotic efficacy and improving therapeutic outcomes in the era of multidrug resistance53-57.
References
1. Mushtaq. Bacterial introduction. 2017. Researchgate.net. Accessed April 24, 2023
2. Tillotson G, Lodise T, Classi P, Mildvan D, McKinnell JA. Antibiotic treatment failure and associated outcomes among adult patients with community-acquired pneu- monia in the outpatient setting: A real-world US insurance claims database study. Open Forum Infect Dis 2020;7: ofaa065
Google Scholar
3. Quinn R. Rethinking antibiotic research and development: World War II and the penicillin collaborative. Am J Public Health 2013;103:426-434
Google Scholar
4. Hobby GL, Meyer K, Chaffee E. Observations on the mechanism of action of penicillin. Proc Soc Exp Biol Med 1942;50:281-285
Google Scholar
5. Bigger J. Treatment of staphylococcal infections with penicillin by intermittent sterilisation. Lancet 1944;244: 497-500
Google Scholar
6. Brauner A, Fridman O, Gefen O, Balaban NQ. Distin- guishing between resistance, tolerance, and persistence to antibiotic treatment. Nat Rev Microbiol 2016;14:320-330
Google Scholar
7. Balaban NQ, Helaine S, Lewis K, Ackermann M, Aldridge B, Andersson DI, et al. Definitions and guidelines for research on antibiotic persistence. Nat Rev Microbiol 2019;17:441-448
Google Scholar
8. Meylan S, Andrews IW, Collins JJ. Targeting antibiotic tolerance, pathogen by pathogen. Cell 2018;172:1228-1238
Google Scholar
9. Levin BR, Concepción-Acevedo J, Udekwu KI. Persistence: A copacetic and parsimonious hypothesis for the existence of non-inherited resistance to antibiotics. Curr Opin Microbiol 2014;21:18-21
Google Scholar
10. Michiels JE, Van den Bergh B, Verstraeten N, Michiels J. Molecular mechanisms and clinical implications of bac- terial persistence. Drug Resist Updat 2016;29:76-89
Google Scholar
11. Radzikowski JL, Schramke H, Heinemann M. Bacterial persistence from a system-level perspective. Curr Opin Biotechnol 2017;46:98-105
Google Scholar
12. Balaban NQ, Merrin J, Chait R, Kowalik L, Leibler S. Bacterial persistence as a phenotypic switch. Science 2004;305:1622-1625
Google Scholar
13. Jõers A, Kaldalu N, Tenson T. The frequency of persisters in Escherichia coli reflects the kinetics of awakening from dormancy. J Bacteriol 2010;192:3379-3384
Google Scholar
14. Levin-Reisman I, Gefen O, Fridman O, Ronin I, Shwa D, Sheftel H, et al. Automated imaging with ScanLag reveals previously undetectable bacterial growth phenotypes. Nat Methods 2010;7:737-739
Google Scholar
15. Manina G, Dhar N, McKinney JD. Stress and host im- munity amplify Mycobacterium tuberculosis phenotypic heterogeneity and induce nongrowing metabolically active forms. Cell Host Microbe 2015;17:32-46
Google Scholar
16. Helaine S, Cheverton AM, Watson KG, Faure LM, Matthews SA, Holden DW. Internalization of Salmonella by macrophages induces the formation of nonreplicating persisters. Science 2014;343:204-208
Google Scholar
17. Luidalepp H, Jõers A, Kaldalu N, Tenson T. Age of in- oculum strongly influences persister frequency and can mask effects of mutations implicated in altered persistence. J Bacteriol 2011;193:3598-3605
Google Scholar
18. Keren I, Kaldalu N, Spoering A, Wang Y, Lewis K. Persister cells and tolerance to antimicrobials. FEMS Microbiol Lett 2004;230:13-18
Google Scholar
19. Akerlund T, Nordström K, Bernander R. Analysis of cell size and DNA content in exponentially growing and stationary-phase batch cultures of Escherichia coli. J Bacteriol 1995;177:6791-6797
Google Scholar
20. Wakamoto Y, Dhar N, Chait R, Schneider K, Signorino-Gelo F, Leibler S, et al. Dynamic persistence of antibiotic-stressed mycobacteria. Science 2013;339:91-95
Google Scholar
21. Maglica Ž, Özdemir E, McKinney JD. Single-cell tracking reveals antibiotic-induced changes in mycobacterial energy metabolism. MBio 2015;6:e02236-14
Google Scholar
22. Dörr T, Vulić M, Lewis K. Ciprofloxacin causes persister formation by inducing the TisB toxin in Escherichia coli. PLoS Biol 2010;8:e1000317
Google Scholar
23. Taylor PC, Schoenknecht FD, Sherris JC, Linner EC. Determination of minimum bactericidal concentrations of oxacillin for Staphylococcus aureus: Influence and significance of technical factors. Antimicrob Agents Chemother 1983;23:142-150
Google Scholar
24. Wang M, Ateia M, Awfa D, Yoshimura C. Regrowth of bacteria after light-based disinfection - What we know and where we go from here. Chemosphere 2021;268: 128850
Google Scholar
25. Dorr T, Lewis K, Vulić M. SOS response induces persist- ence to fluoroquinolones in Escherichia coli. PLoS Genet 2009;5:e1000760
Google Scholar
26. 26. Kim Y, Wood TK. Toxins Hha and CspD and small RNA regulator Hfq are involved in persister cell formation through MqsR in Escherichia coli. Biochem Biophys Res Commun 2010;391:209-213
Google Scholar
27. Johnson PJ, Levin BR. Pharmacodynamics, population dynamics, and the evolution of persistence in Staphylo- coccus aureus. PLoS Genet 2013;9:e1003123
Google Scholar
28. Luria SE, Delbrück M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 1943;28:491-511
Google Scholar
29. Nierman WC, Yu Y, Losada L. The in vitro antibiotic-tolerant persister population in Burkholderia pseudomallei is altered by environmental factors. Front Microbiol 2015; 6:1338
Google Scholar
30. Gefen O, Gabay C, Mumcuoglu M, Engel G, Balaban NQ. Single-cell protein induction dynamics reveals a period of vulnerability to antibiotics in persister bacteria. Proc Natl Acad Sci U S A 2008;105:6145-6149
Google Scholar
31. Goneau LW, Yeoh NS, MacDonald KW, Cadieux PA, Burton JP, Razvi H, et al. Selective target inactivation rather than global metabolic dormancy causes antibiotic tolerance in uropathogens. Antimicrob Agents Chemother 2014;58:2089-2097
Google Scholar
32. Kussell E, Leibler S. Phenotypic diversity, population growth, and information in fluctuating environments. Science 2005;309:2075-2078
Google Scholar
33. Kotte O, Volkmer B, Radzikowski JL, Heinemann M. Phenotypic bistability in Escherichia coli's central carbon metabolism. Mol Syst Biol 2014;10:736
Google Scholar
34. Spoering AL, Vulić M, Lewis K. GlpD and PlsB participate in persister cell formation in Escherichia coli. J Bacteriol 2006;188:5136-5144
Google Scholar
35. Hansen S, Lewis K, Vulić M. Role of global regulators and nucleotide metabolism in antibiotic tolerance in Escherichia coli. Antimicrob Agents Chemother 2008;52: 2718-2726
Google Scholar
36. Bojsen R, Regenberg B, Gresham D, Folkesson A. A common mechanism involving the TORC1 pathway can lead to amphotericin B persistence in biofilm and plank- tonic Saccharomyces cerevisiae populations. Sci Rep 2016;6:21874
Google Scholar
37. Henry TC, Brynildsen MP. Development of Persister-FACSeq: A method to massively parallelize quantification of persister physiology and its heterogeneity. Sci Rep 2016;6:25100
Google Scholar
38. Vazquez-Laslop N, Lee H, Neyfakh AA. Increased persist- ence in Escherichia coli caused by controlled expression of toxins or other unrelated proteins. J Bacteriol 2006; 188:3494-3497
Google Scholar
39. 39. Moyed HS, Bertrand KP. hipA, a newly recognized gene of Escherichia coli K-12 that affects the frequency of persistence after inhibition of murein synthesis. J Bacteriol 1983;155:768-775
Google Scholar
40. Shah D, Zhang Z, Khodursky A, Kaldalu N, Kurg K, Lewis K. Persisters: A distinct physiological state of E. coli. BMC Microbiol 2006;6:53
Google Scholar
41. Lewis K. Persister cells. Annu Rev Microbiol 2010;64:357-372
Google Scholar
42. Sugimoto S, Arita-Morioka KI, Mizunoe Y, Yamanaka K, Ogura T. Thioflavin T as a fluorescence probe for moni- toring RNA metabolism at molecular and cellular levels. Nucleic Acids Res 2015;43:e92
Google Scholar
43. Rice LB. Progress and challenges in implementing the research on ESKAPE pathogens. Infect Control Hosp Epidemiol 2010;31 Suppl 1:S7-S10
Google Scholar
44. Tacconelli E, Carrara E, Savoldi A, Harbarth S, Mendelson M, Monnet DL, et al. Discovery, research, and devel- opment of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis 2018;18:318-327
Google Scholar
45. Elsner HA, Sobottka I, Mack D, Claussen M, Laufs R, Wirth R. Virulence factors of Enterococcus faecalis and Enterococcus faecium blood culture isolates. Eur J Clin Microbiol Infect Dis 2000;19:39-42
Google Scholar
46. ECDC. Surveillance of antimicrobial resistance in Europe 2017. European Centre for Disease Prevention and Control.
47. Solomon SL, Oliver KB. Antibiotic resistance threats in the United States: Stepping back from the brink. Am Fam Physician 2014;89:938-941
Google Scholar
48. Lowy FD. Staphylococcus aureus infections. N Engl J Med 1998;339:520-532
Google Scholar
49. Skinner D, Keefer CS. Significance of bacteremia caused by Staphylococcus aureus: A study of one hundred and twenty-two cases and a review of the literature concerned with experimental infection in animals. Arch Intern Med 1941;68:851-875
Google Scholar
50. Brown PD, Ngeno C. Antimicrobial resistance in clinical isolates of Staphylococcus aureus from hospital and community sources in southern Jamaica. Int J Infect Dis 2007;11:220-225
Google Scholar
51. Lacey RW. Mechanisms of resistance to beta-lactam antibiotics in Staphylococcus aureus. Scand J Infect Dis Suppl 1984;42:64-71
Google Scholar
52. 52. Harkins CP, Pichon B, Doumith M, Parkhill J, Westh H, Tomasz A, et al. Methicillin-resistant Staphylococcus aureus emerged long before the introduction of methi- cillin into clinical practice. Genome Biol 2017;18:130
Google Scholar
53. Hassoun A, Linden PK, Friedman B. Incidence, prevalence, and management of MRSA bacteremia across patient populations: A review of recent developments in MRSA management and treatment. Crit Care 2017;21:211
Google Scholar
54. Hartman BJ, Tomasz A. Low-affinity penicillin-binding protein associated with beta-lactam resistance in Staphylo- coccus aureus. J Bacteriol 1984;158:513-516
Google Scholar
55. Tenover FC, Goering RV. Methicillin-resistant Staphylo- coccus aureus strain USA300: Origin and epidemiology. J Antimicrob Chemother 2009;64:441-446
Google Scholar
56. Benoit JB, Frank DN, Bessesen MT. Genomic evolution of Staphylococcus aureus isolates colonizing the nares and progressing to bacteremia. PLoS One 2018;13:e0195860
Google Scholar
57. Aarian T, Daum RS, Petty LA, Steinbeck JL, Yin Z, Nolan D, et al. Intrahost evolution of methicillin-resistant Staphylococcus aureus USA300 among individuals with recurring skin and soft-tissue infections. J Infect Dis 2016; 214:895-905
Google Scholar
Congratulatory MessageClick here!